Protein Function and Molecular Interactions – Exploring the Molecular Machinery of Life
Proteins are the dynamic molecular machines that drive virtually every process in living cells from catalyzing reactions and transmitting signals to providing structural support and regulating gene expression. Their function emerges directly from their intricate three-dimensional structures and sophisticated interaction networks with ligands, other proteins, nucleic acids, and cofactors. Understanding these mechanisms is foundational to drug discovery, synthetic biology, personalized medicine, and biotechnology innovation.
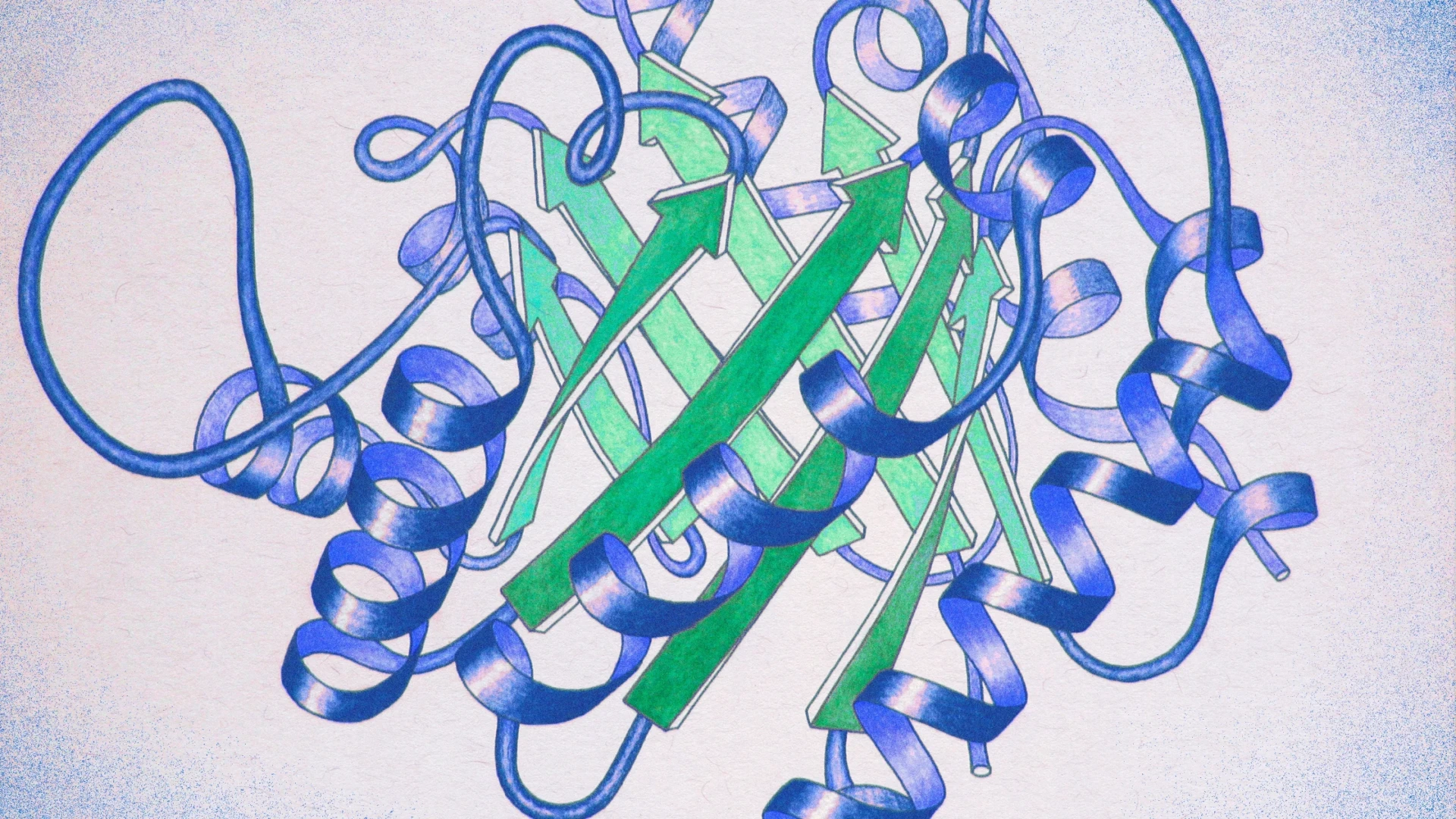
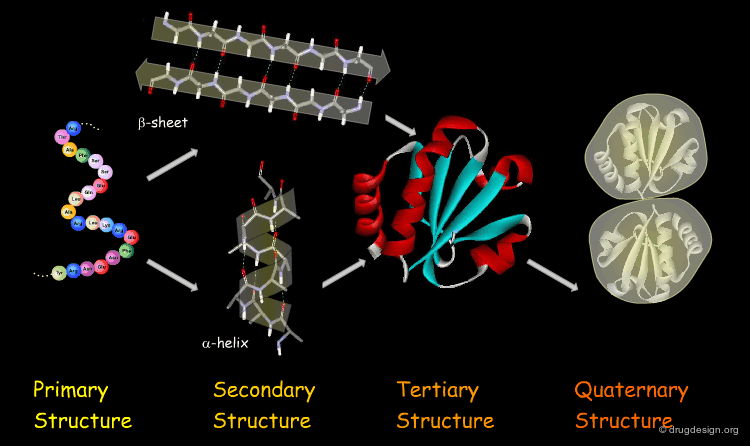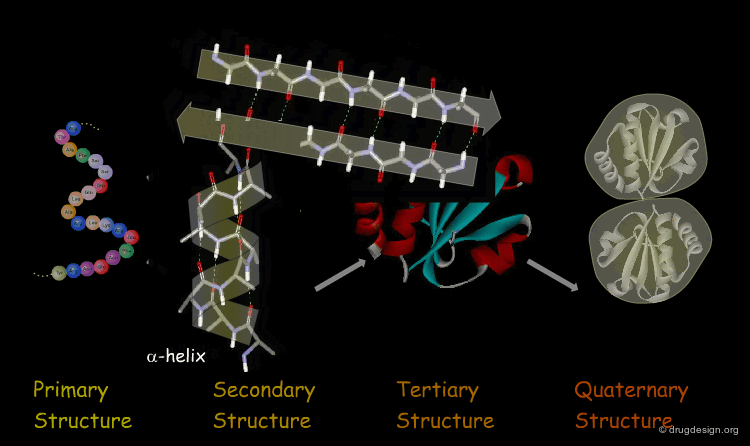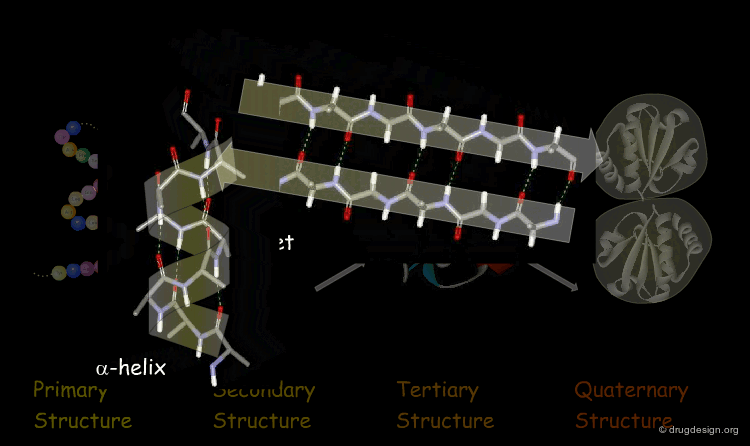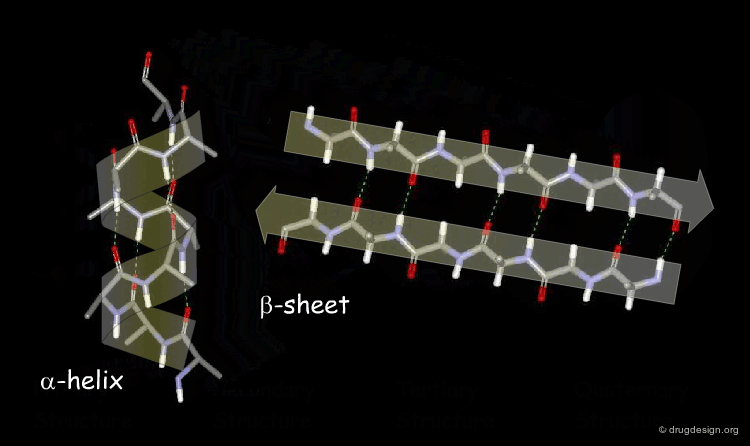
1. Protein Structure – The Foundation of Function
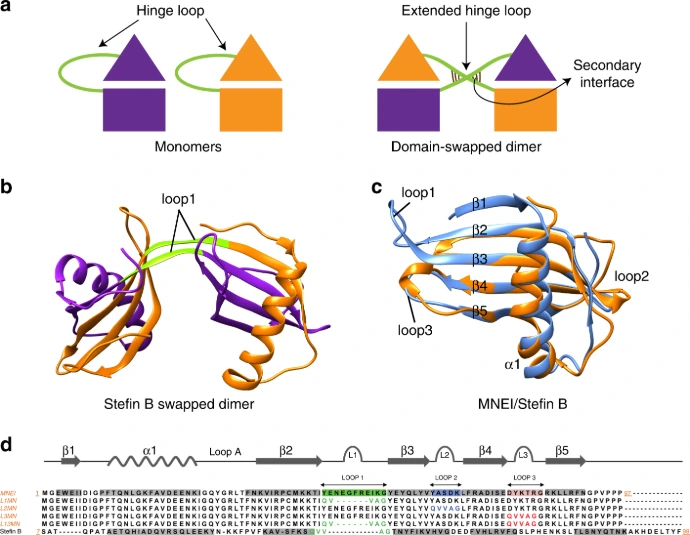
Proteins fold into precise three-dimensional architectures determined by their amino acid sequence, governed by physicochemical principles including hydrophobic collapse, hydrogen bonding, electrostatic interactions, and van der Waals forces. Primary Structure: The linear sequence of amino acids encoded by genes.
Secondary Structure: Local folding into α-helices, β-sheets, and turns/loops stabilized by backbone hydrogen bonds.
Tertiary Structure: Overall 3D fold bringing distant residues together to form functional sites (active sites, binding pockets).
Quaternary Structure: Assembly of multiple polypeptide chains into oligomeric complexes, often essential for activity (e.g., hemoglobin tetramer).
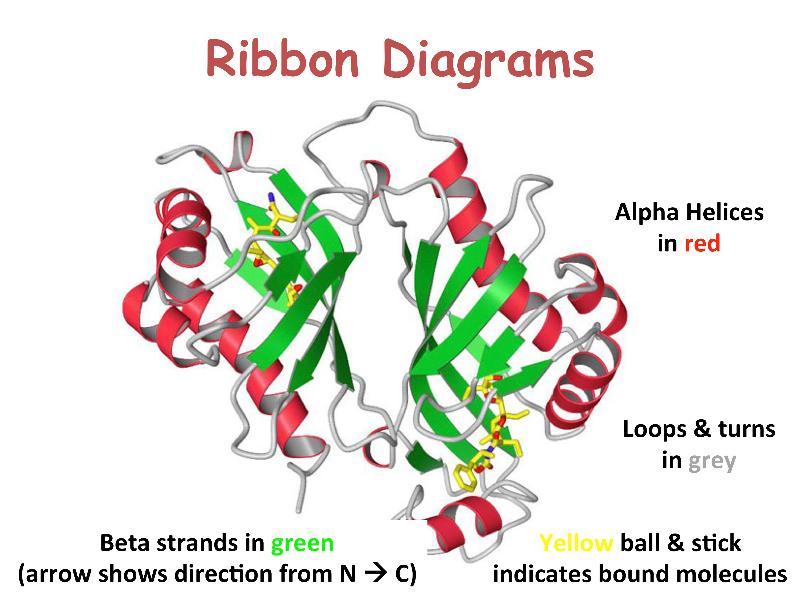
2 . Protein-Ligand Interactions
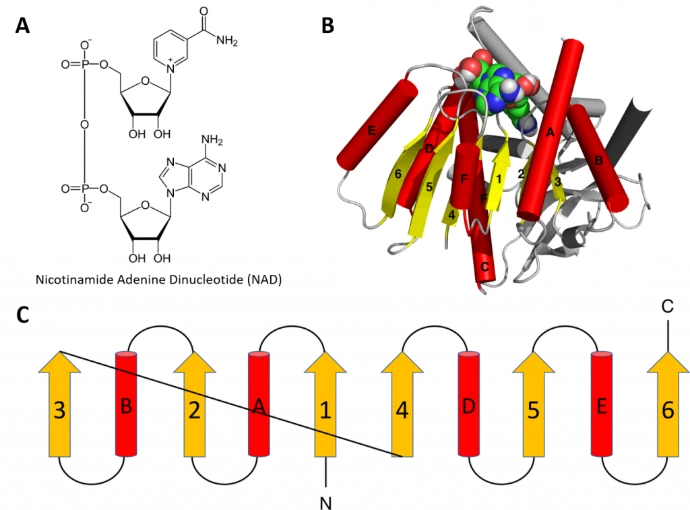
Protein function often depends on precise, reversible binding of small molecules (substrates, cofactors, allosteric modulators)
Binding Affinity (K_d): Quantified by dissociation constant; tighter binding (lower K_d) enhances efficiency.
Specificity: Shape and chemical complementarity (lock-and-key or induced-fit models) ensure selectivity.
Allosteric Regulation: Binding at a distant (allosteric) site modulates activity at the orthosteric site through conformational changes or dynamic shifts.
Did you know?
The human interactome is estimated to contain ~150,000–300,000 binary interactions, many still structurally unresolved.
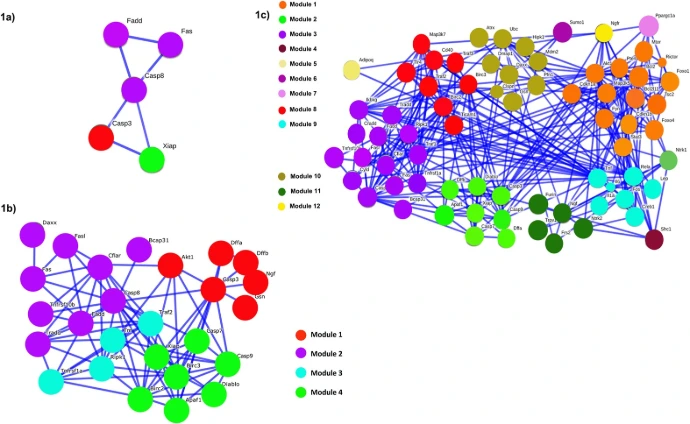
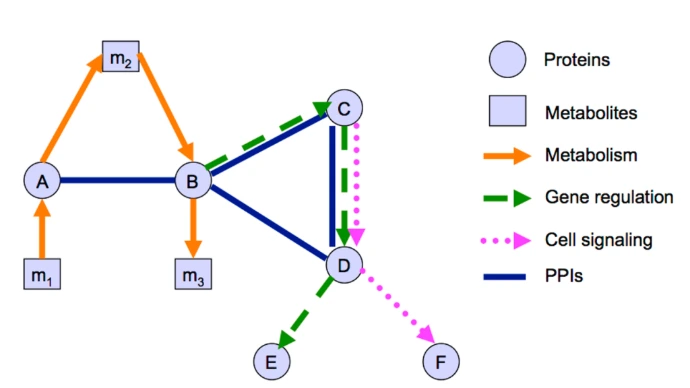
3. Protein-Protein Networks
Proteins operate within dense interaction networks (interactomes) that integrate signals, coordinate metabolism, and maintain cellular architecture.
- Signal Transduction: Cascades (e.g., MAPK pathway) transmit external cues to internal responses.
- Metabolic Pathways: Multi-enzyme complexes channel intermediates efficiently (substrate channeling).
- Structural Assemblies: Cytoskeleton (actin, microtubules) and complexes like ribosomes or proteasomes.
4. Functional Domains and Motifs
Modular architecture allows proteins to combine reusable functional units.
- Enzymatic Domains: e.g., kinase, protease, or glycosyltransferase domains.
- Binding Domains: SH2/SH3 (phosphotyrosine/ proline-rich), PDZ (C-terminal motifs).
- Regulatory Motifs: Nuclear localization signals, degradation tags (e.g., PEST sequences).
5. Proteomics – Large-Scale Protein Analysis
Proteomics maps the proteome dynamically:
- Mass Spectrometry (MS): Identifies proteins, PTMs (phosphorylation, ubiquitination), and quantifies changes (TMT, label-free).
- Affinity Proteomics: Antibody/protein arrays detect interactions.
- Structural Proteomics: Cross-linking MS reveals interaction interfaces.
Conclusion
Proteins embody life's molecular ingenuity their structures dictate function, interactions orchestrate complexity, and engineering unlocks innovation. From fundamental mechanisms to translational breakthroughs, protein science remains at the heart of biotechnology and medicine.
Explore these concepts further in BioStruct Explorer's curated datasets, interactive 3D viewers, and network tools.