Protein Conformational Dynamics and Binding Site Flexibility
Introduction
Proteins are highly dynamic macromolecules. Their conformational flexibility is crucial for molecular recognition, ligand binding specificity, and biological function. While static structures (from X-ray, cryo-EM, etc.) offer valuable snapshots, they miss the full spectrum of motions essential for understanding binding mechanisms and predicting interactions.
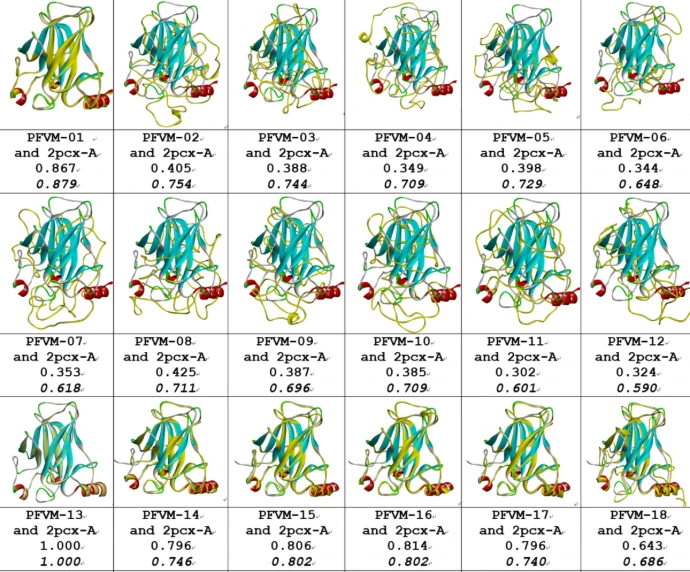
Conformational Ensembles
Proteins in solution exist as dynamic ensembles of interconverting conformations rather than a single rigid structure.
- Sub-populations within the ensemble can preferentially bind different ligands.
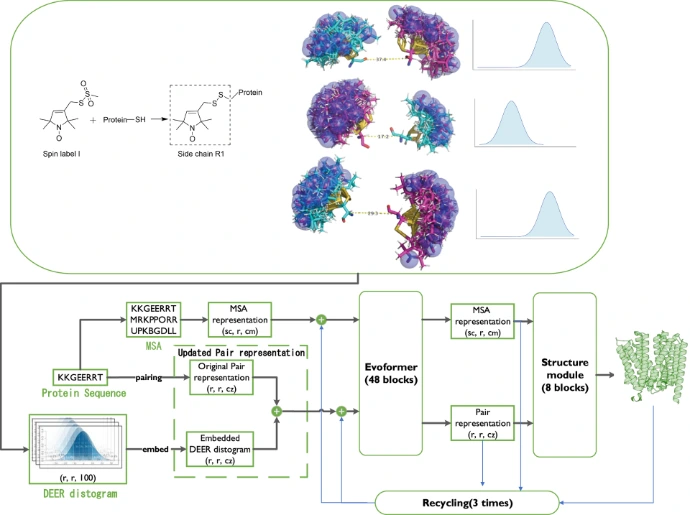
- Key experimental and computational methods include NMR spectroscopy, crystallographic ensembles, and molecular dynamics (MD) simulations to map these states.
Molecular Recognition Hotspots – Anchors of Protein-Ligand Interactions
Molecular recognition hotspots are key residues that stabilize ligand binding.
They act as anchors, defining specificity and interaction patterns.
Mapping hotspots reveals conserved motifs across protein families.
1. What Are Hotspots?
Hotspots contribute disproportionately to binding energy.
They include aromatic, charged, and hydrogen-bonding residues.
Even a few residues can control the stability and specificity of the complex.

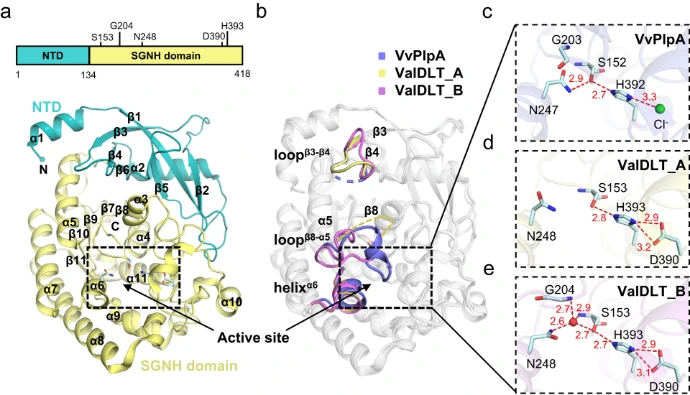
3. Molecular Recognition Hotspots
Certain "hotspot" residues serve as primary anchors for interactions.
- Mapping these hotspots predicts binding patterns and ligand specificity.
- Cross-family comparisons uncover conserved motifs essential for recognition.
2. Mapping Hotspots
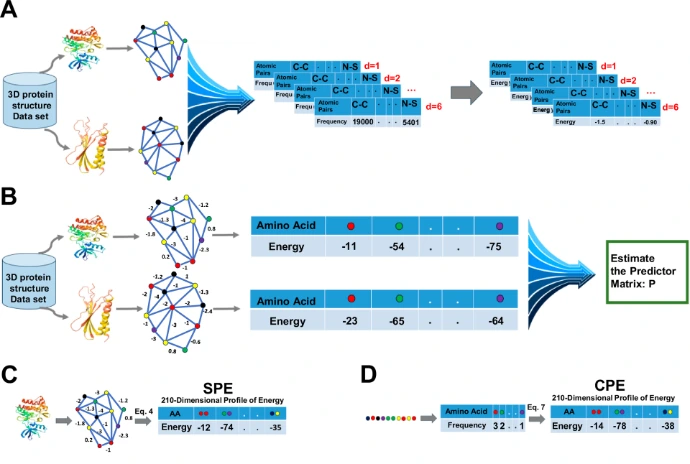
Hotspots are identified using:
3D structural analysis
Computational energy scoring
Cross-family comparison
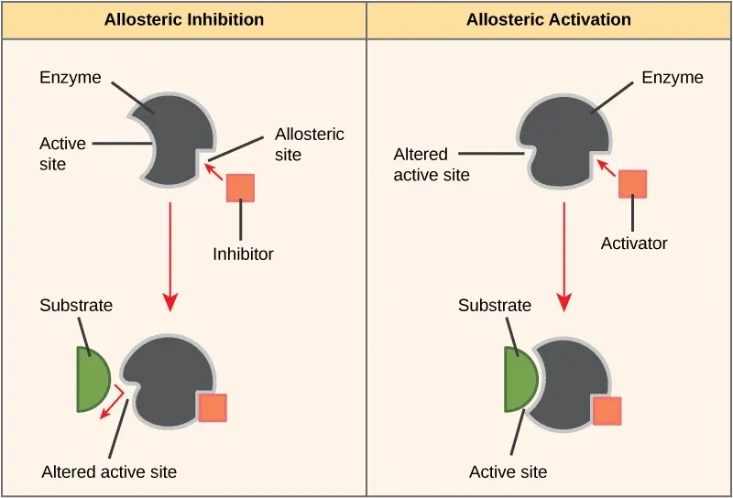
Allosteric Modulation and Functional Implications
Ligand binding at distant (allosteric) sites can propagate conformational changes to the active site.
- Allosteric sites act as regulatory switches in enzymes and signaling pathways.
- Structural studies help identify functional switches across protein families.
Computational Approaches
- Standard MD simulations track transitions over time.
- Enhanced sampling (e.g., replica-exchange MD, metadynamics) captures rare, functionally important states.
- Incorporating structural ensembles into docking and bioinformatics pipelines boosts prediction accuracy.
Structural Flexibility Across Protein Families
Comparative analyses reveal families with high vs. low flexibility.
- Higher flexibility often correlates with ligand diversity, catalytic promiscuity, and regulatory versatility.
- Structural classification guides prioritization of experimental targets.