Comparative Analysis of Protein Binding Sites Across Biological Systems
1. Introduction
Comparative analysis of protein binding sites across diverse biological systems ranging from bacteria to humans and across homologous protein families reveals key evolutionary patterns, degrees of functional conservation, and underlying molecular "design rules" that govern ligand recognition. By structurally aligning binding pockets (e.g., using tools like TM-align or PocketMatch), researchers can map conserved interactions (e.g., hydrogen bonds, hydrophobic contacts) and predict cross-species ligand compatibility, aiding drug repurposing, evolutionary studies, and functional annotation of novel proteins.
Key benefits include identifying selective pressures on functional regions and detecting adaptations that lead to specificity or new functions.
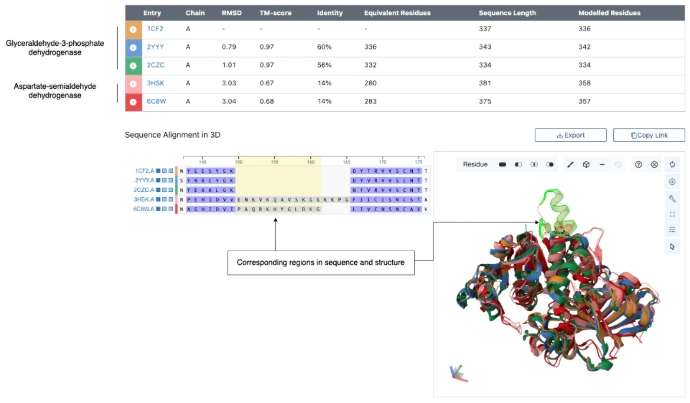
2. Classification of Binding Sites
Protein binding sites, or pockets, are classified according to their structural and chemical features.
Geometry and shape determine how a ligand fits within the cavity. Volume, depth, and surface curvature influence binding stability and specificity.
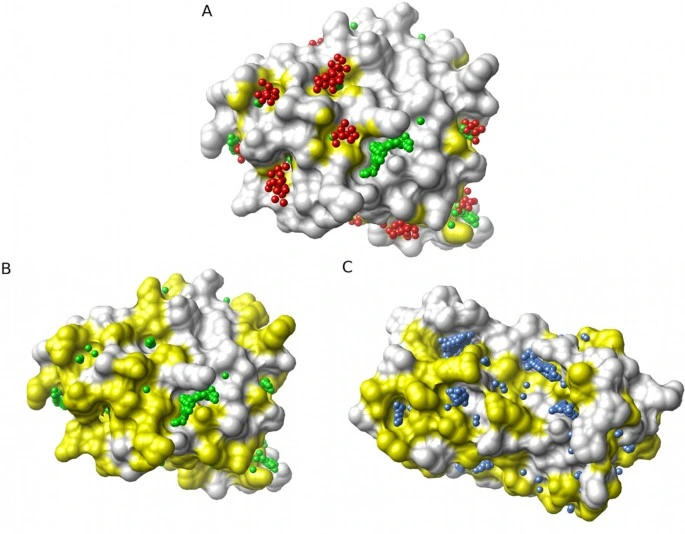
Chemical properties define interaction potential. Hydrophobic regions, charged residues, and hydrogen-bonding groups shape ligand orientation and complementarity.
Residue conservation highlights functional importance. Highly conserved amino acids often play key roles in binding and structural stability.
Binding sites may be canonical, representing common and well-defined pockets, or non-canonical, including shallow or allosteric regions that require detailed structural analysis for proper annotation.
3. Conserved vs. Variable Sites
- Conserved sites/residues maintain core functions (e.g., catalytic triads in enzymes or key interaction motifs), under strong purifying selection across distant species.
- Variable sites allow specificity, environmental adaptation, or evolutionary innovation (e.g., altered ligand affinity in orthologs).
- Patterns from comparative studies inform predictive models, such as machine learning for binding affinity or allosteric regulation.
4. Ligand Diversity Across Homologs
Homologs frequently bind chemically related ligands but exhibit affinity variations due to subtle structural tweaks.
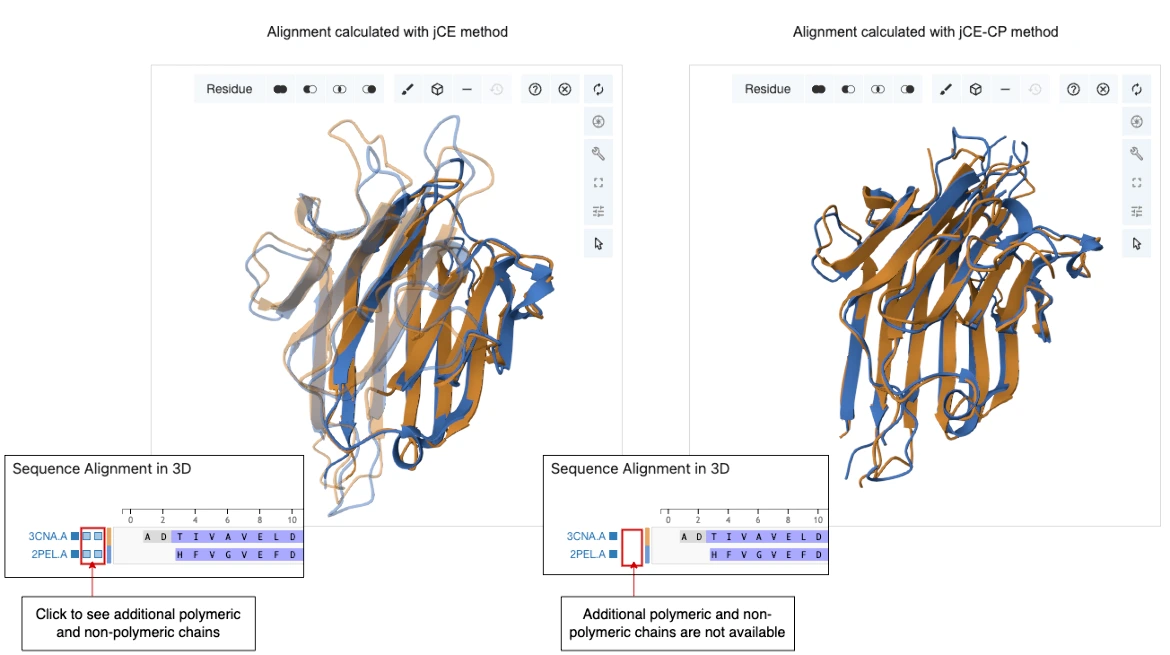
- Structural mapping (e.g., via superposition) distinguishes shared core interactions from species-specific ones.
- This reveals functional redundancy (e.g., broad substrate acceptance) vs. specialization (e.g., drug resistance mutations).
5. Evolutionary Insights
High structural conservation in binding sites indicates strong selective pressure for function preservation.
- Variations often signal adaptive evolution (e.g., new ligand recognition) or sub-functionalization.
- Comparative analyses accelerate functional annotation of uncharacterized proteins in understudied genomes.
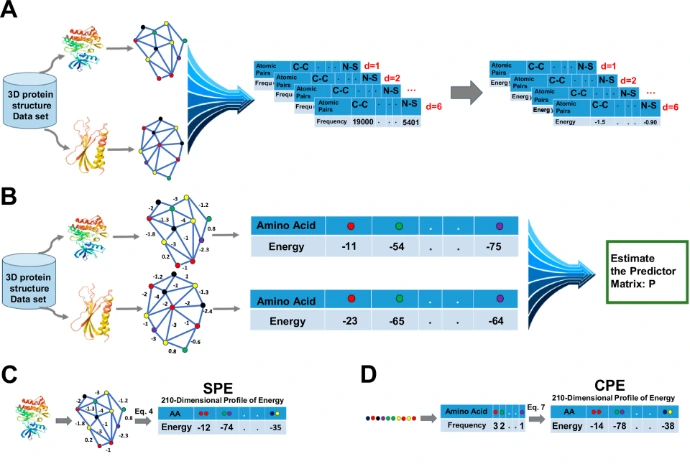
Recent methods use energy profiles for rapid cross-species comparison.
6. Integrating Structural and Sequence Data
Combining 3D alignments (e.g., TM-align) with sequence conservation (e.g., multiple sequence alignments) enhances binding site prediction accuracy.
- Multi-species datasets highlight critical motifs.
- Supports reliable cross-species inference in protein families.

7. Applications in Research
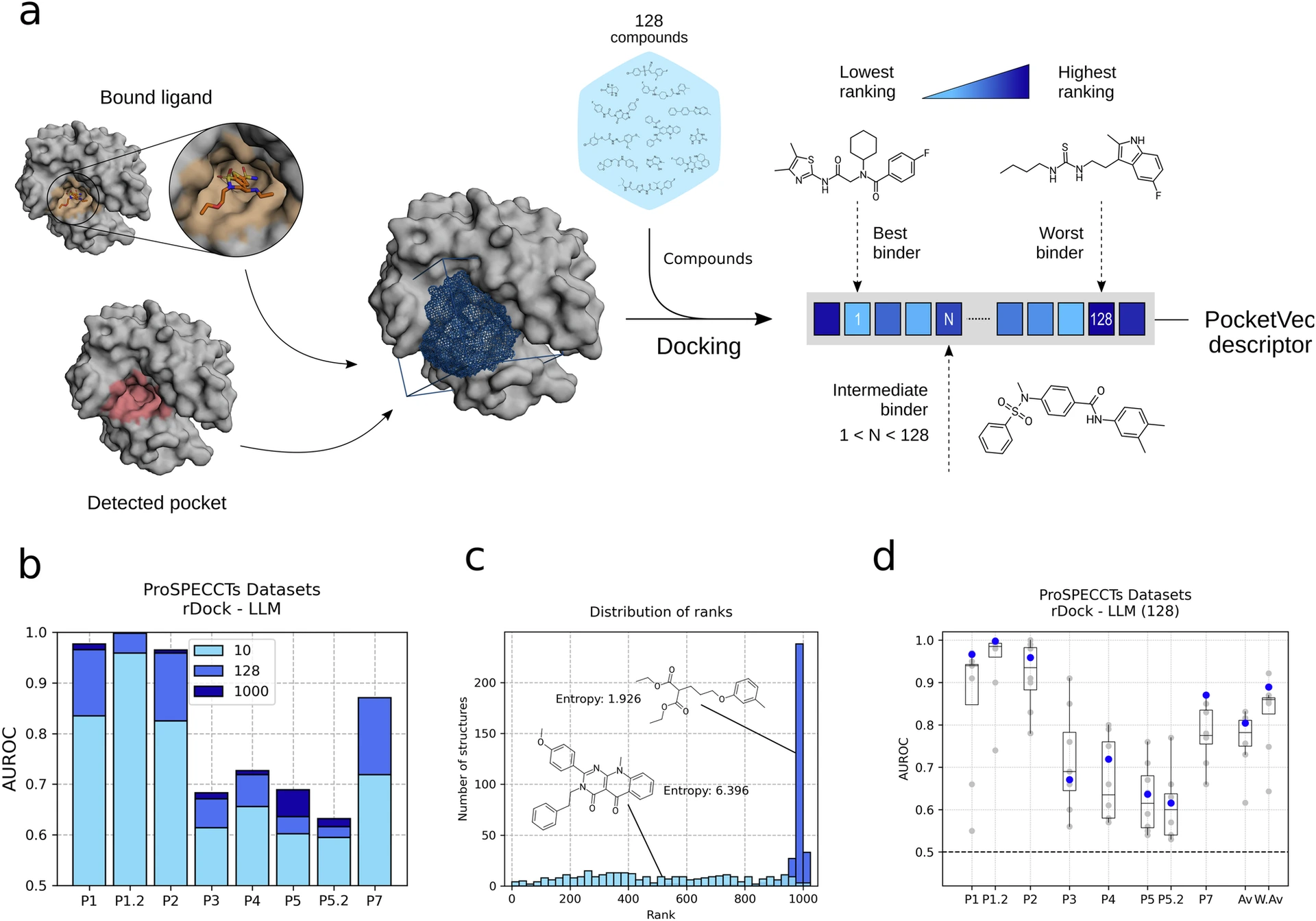
- Benchmarking/improving binding site prediction algorithms (e.g., comparative evaluation of tools).
- Mapping protein-ligand interaction networks across proteomes.
- Guiding synthetic biology by identifying transplantable conserved motifs.
8. Future Directions
Incorporate protein dynamics (e.g., conformational ensembles from MD simulations) for more realistic comparisons.
- Leverage AI/ML (e.g., deep learning for automated pattern recognition in large datasets).
- Expand structural databases (e.g., via AlphaFold) to include understudied organisms, building a comprehensive protein-ligand knowledge base.